This class is derived from chent. It makes it easy to create a chent from the ISO common name of a pesticide active ingredient, and additionally stores the ISO name as well as the complete result of querying the BCPC compendium using bcpc_query.
Format
An R6::R6Class generator object
Super class
chents::chent -> pai
Public fields
isoISO common name of the active ingredient according to ISO 1750
bcpcInformation retrieved from the BCPC compendium available online at <pesticidecompendium.bcpc.org>
Methods
Inherited methods
chents::chent$add_PUF()chents::chent$add_TP()chents::chent$add_cwsat()chents::chent$add_p0()chents::chent$add_soil_degradation()chents::chent$add_soil_ff()chents::chent$add_soil_sorption()chents::chent$add_transformation()chents::chent$emf()chents::chent$get_chyaml()chents::chent$get_pubchem()chents::chent$get_rdkit()chents::chent$pdf()chents::chent$png()chents::chent$try_pubchem()
Method new()
Create a new pai object
Usage
pai$new(
iso,
identifier = iso,
smiles = NULL,
inchikey = NULL,
bcpc = TRUE,
pubchem = TRUE,
pubchem_from = "auto",
rdkit = TRUE,
template = NULL,
chyaml = FALSE
)Arguments
isoThe ISO common name to be used in the query of the BCPC compendium
identifierAlternative identifier used for querying pubchem
smilesOptional user provided SMILES code
inchikeyOptional user provided InChI Key
bcpcShould the BCPC compendium be queried?
pubchemShould an attempt be made to retrieve chemical information from PubChem via the webchem package?
pubchem_fromPossibility to select the argument that is used to query pubchem
rdkitShould an attempt be made to retrieve chemical information from a local rdkit installation via python and the reticulate package?
templateAn optional SMILES code to be used as template for RDKit
chyamlShould we look for a identifier.yaml file in the working
Examples
# Don't run examples per default, as PubChem may be unavailable
# \dontrun{
atr <- pai$new("atrazine")
#> Querying BCPC for atrazine ...
#> Querying PubChem for inchikey MXWJVTOOROXGIU-UHFFFAOYSA-N ...
#> Get chemical information from RDKit using PubChem SMILES
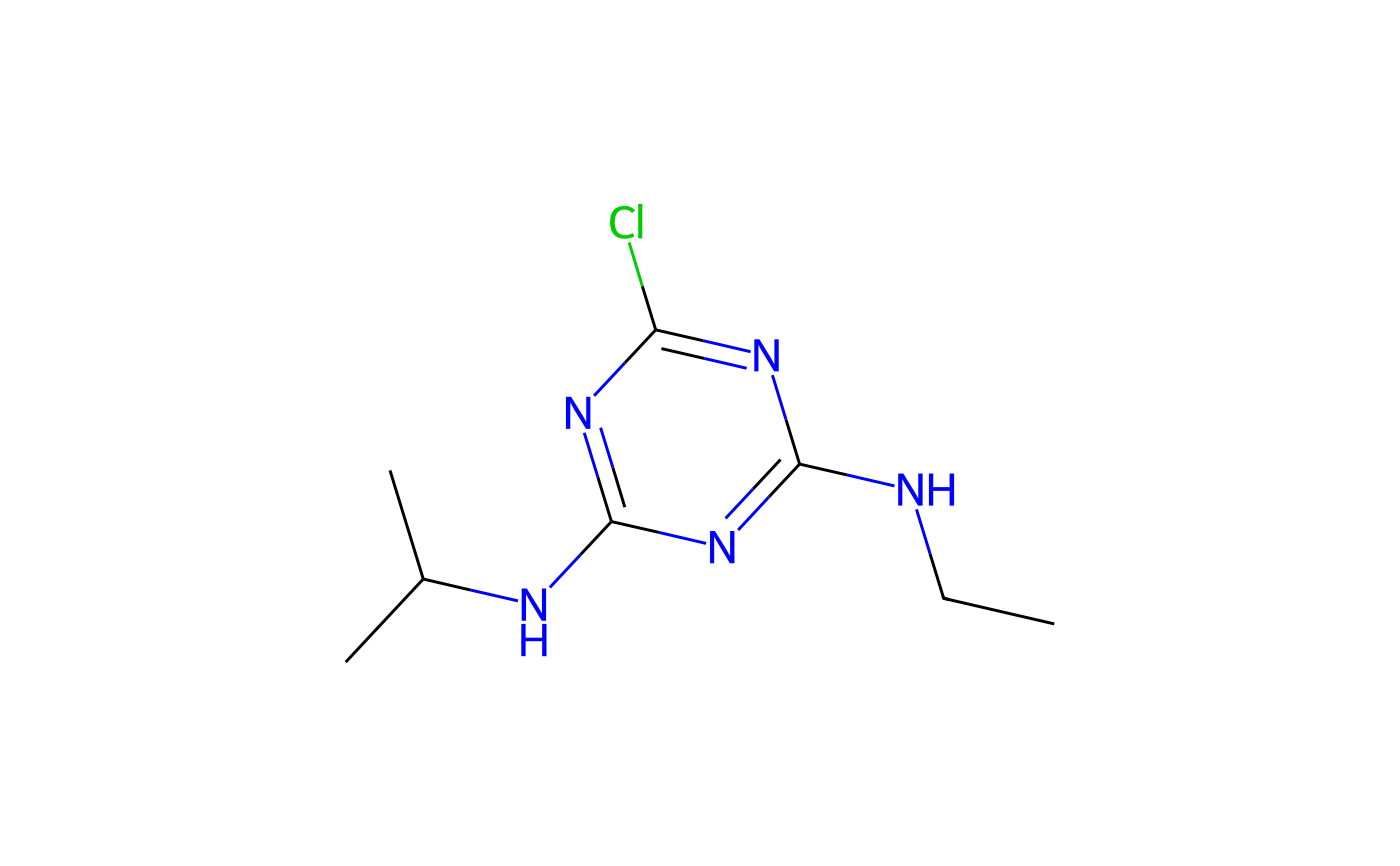
#> CCNC1=NC(=NC(=N1)Cl)NC(C)C
print(atr)
#> <pai> with ISO common name $iso atrazine
#> <chent>
#> Identifier $identifier atrazine
#> InChI Key $inchikey MXWJVTOOROXGIU-UHFFFAOYSA-N
#> SMILES string $smiles:
#> PubChem
#> "CCNC1=NC(=NC(=N1)Cl)NC(C)C"
#> Molecular weight $mw: 215.7
#> PubChem synonyms (up to 10):
#> [1] "atrazine" "1912-24-9" "Gesaprim" "Aatrex" "Atranex"
#> [6] "Atrazin" "Oleogesaprim" "Atazinax" "Atrasine" "Chromozin"
if (!is.null(atr$Picture)) {
plot(atr)
}
 # We can also define pais that are not found on the BCPC site
decanol <- pai$new("1-Decanol")
#> Querying BCPC for 1-Decanol ...
#> Common name 1-Decanol is not known at the BCPC compendium, trying PubChem
#> Querying PubChem for name 1-Decanol ...
#> Get chemical information from RDKit using PubChem SMILES
#> CCCCCCCCCCO
print(decanol)
#> <pai> without ISO common name
#> <chent>
#> Identifier $identifier 1-Decanol
#> InChI Key $inchikey MWKFXSUHUHTGQN-UHFFFAOYSA-N
#> SMILES string $smiles:
#> PubChem
#> "CCCCCCCCCCO"
#> Molecular weight $mw: 158.3
#> PubChem synonyms (up to 10):
#> [1] "1-DECANOL" "Decan-1-ol" "Decyl alcohol" "112-30-1"
#> [5] "n-Decyl alcohol" "n-Decanol" "Capric alcohol" "Nonylcarbinol"
#> [9] "Antak" "Royaltac"
# }
# We can also define pais that are not found on the BCPC site
decanol <- pai$new("1-Decanol")
#> Querying BCPC for 1-Decanol ...
#> Common name 1-Decanol is not known at the BCPC compendium, trying PubChem
#> Querying PubChem for name 1-Decanol ...
#> Get chemical information from RDKit using PubChem SMILES
#> CCCCCCCCCCO
print(decanol)
#> <pai> without ISO common name
#> <chent>
#> Identifier $identifier 1-Decanol
#> InChI Key $inchikey MWKFXSUHUHTGQN-UHFFFAOYSA-N
#> SMILES string $smiles:
#> PubChem
#> "CCCCCCCCCCO"
#> Molecular weight $mw: 158.3
#> PubChem synonyms (up to 10):
#> [1] "1-DECANOL" "Decan-1-ol" "Decyl alcohol" "112-30-1"
#> [5] "n-Decyl alcohol" "n-Decanol" "Capric alcohol" "Nonylcarbinol"
#> [9] "Antak" "Royaltac"
# }